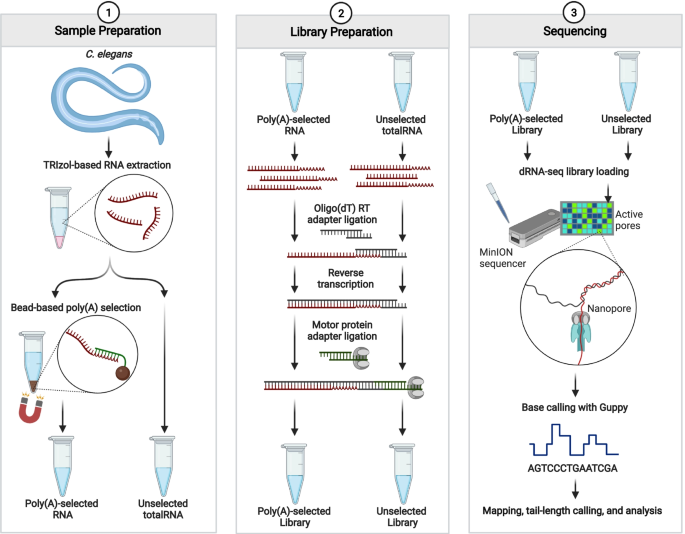
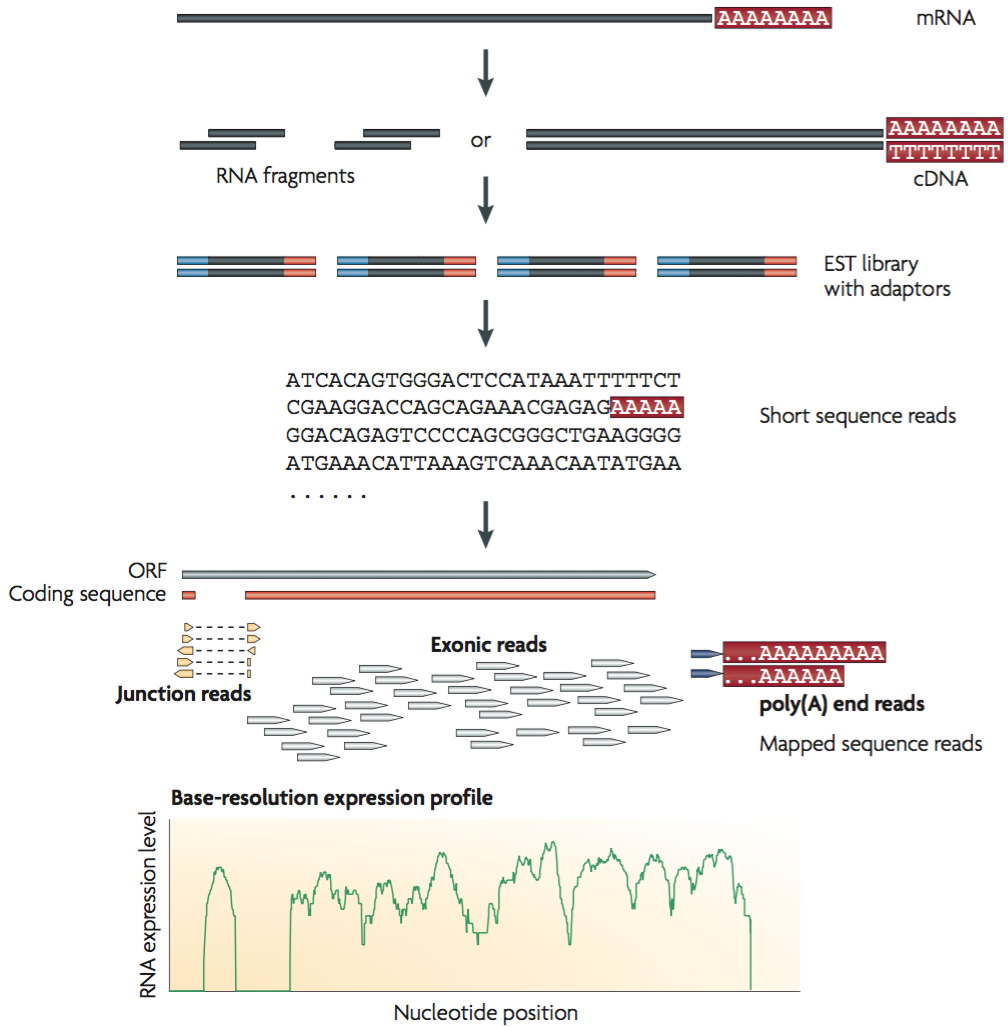
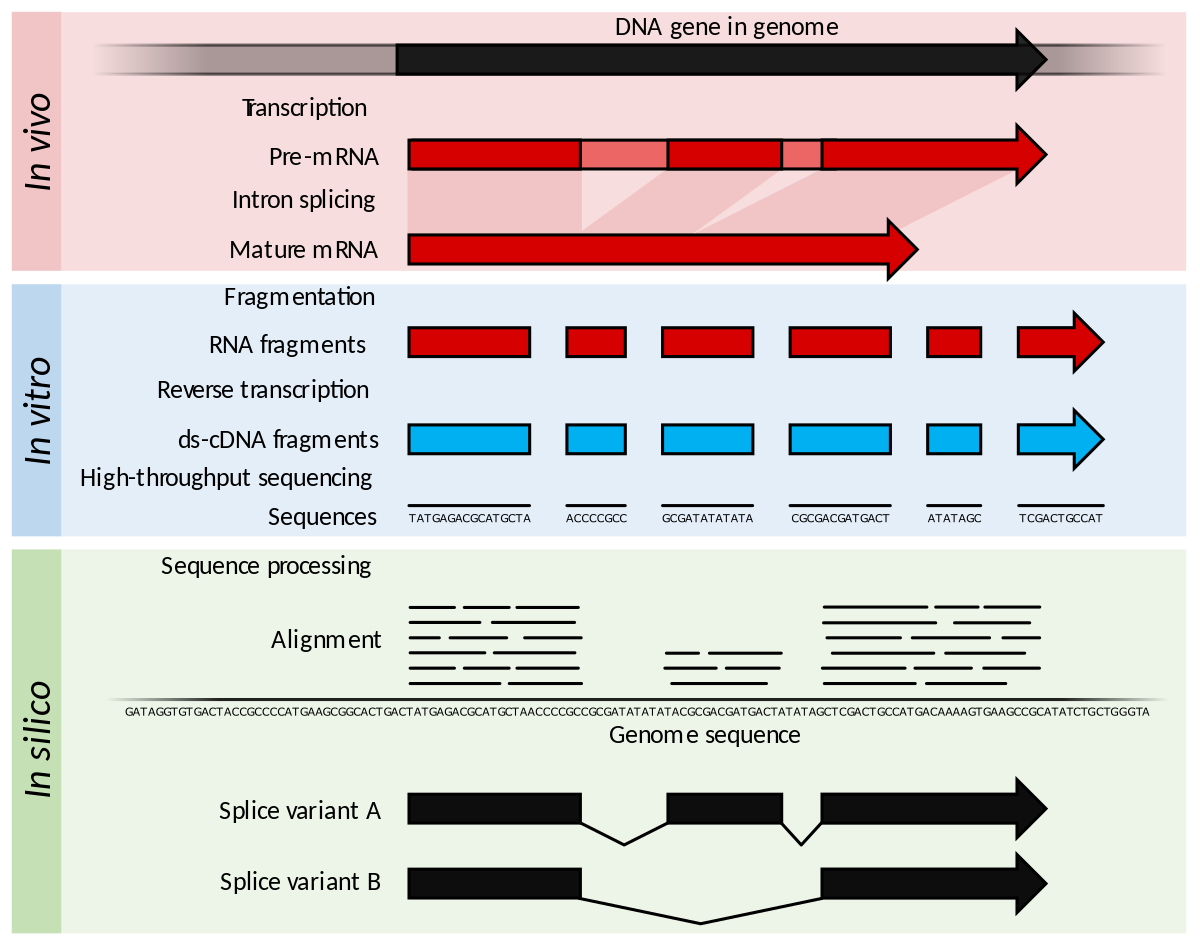
A Rapid, Simple, and Inexpensive Method for the Preparation of Strand-Specific RNA-Seq Libraries - High-throughput sequencing o… | Gene expression, Method, Library

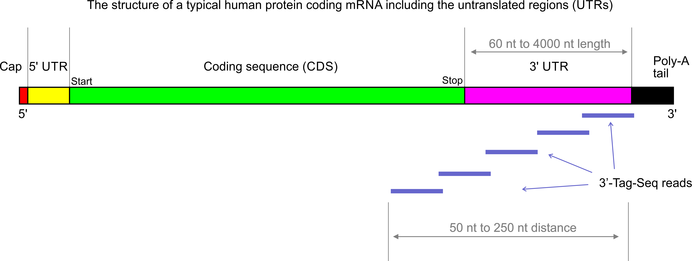
Poly(A)-seq – A method for direct sequencing and analysis of the transcriptomic poly(A)-tails | RNA-Seq Blog

Poly(A) capture full length cDNA sequencing improves the accuracy and detection ability of transcript quantification and alternative splicing events | Scientific Reports

Comparison of RNA-Seq by poly (A) capture, ribosomal RNA depletion, and DNA microarray for expression profiling | BMC Genomics | Full Text

Diagram of LM-Seq sample preparation protocol. Poly-A-tailed mRNA is... | Download Scientific Diagram

A multiplex RNA-seq strategy to profile poly(A+) RNA: Application to analysis of transcription response and 3′ end formation - ScienceDirect
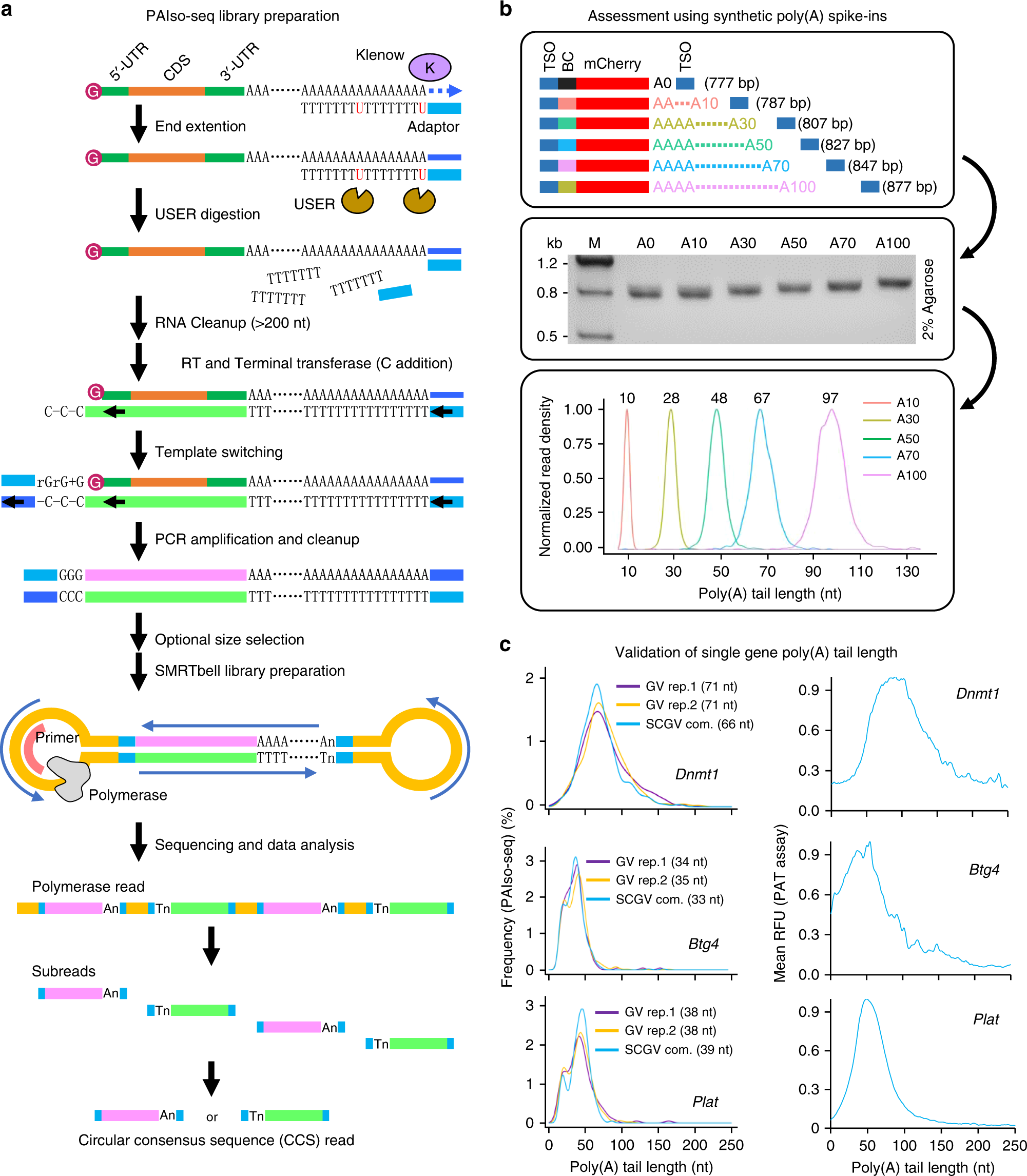
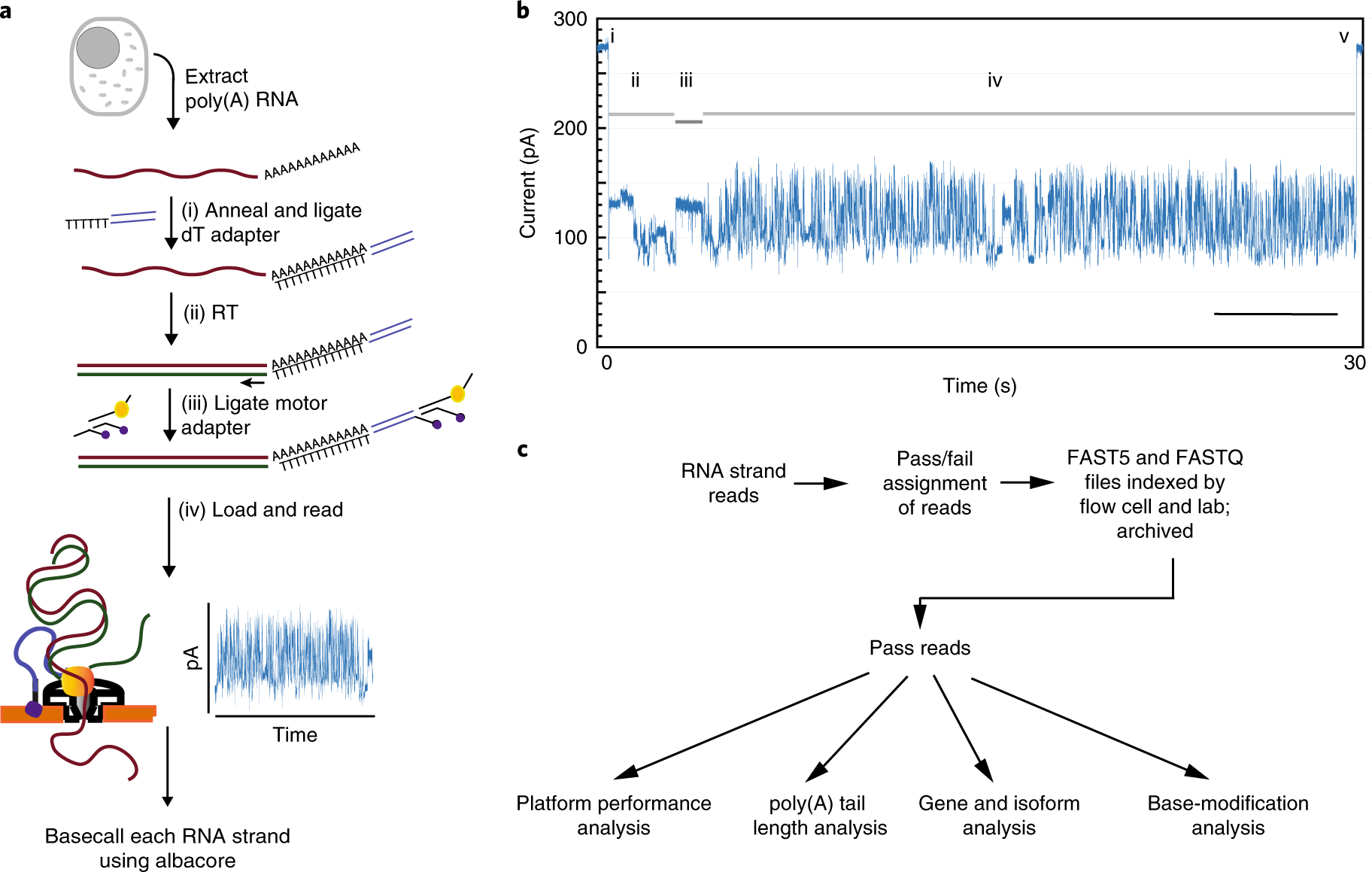
Poly(A) inclusive RNA isoform sequencing (PAIso−seq) reveals wide-spread non-adenosine residues within RNA poly(A) tails | Nature Communications

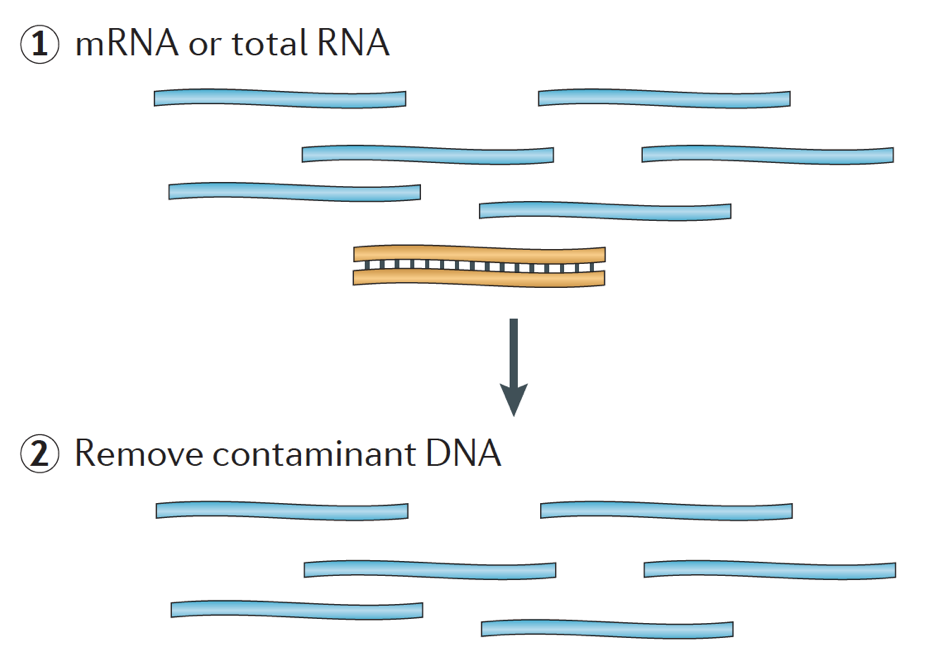
Poly(A)-ClickSeq – click-chemistry for next-generation 3΄-end sequencing without RNA enrichment or fragmentation | RNA-Seq Blog